-
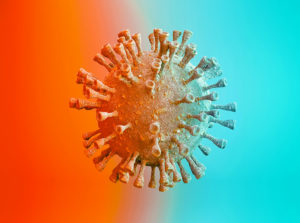
COVID-19
ISB researchers, along with the global scientific community at large, immediately focused their attention on the COVID-19 pandemic and the virus that causes the disease – SARS-CoV-2. ISB President Dr. Jim Heath and scientists across labs and disciplines at ISB have worked to uncover the secrets behind COVID-19.
-
Healthy Aging
Research suggests that it may be possible to eliminate the correlation between age and diseases. ISB is working to leverage systems biology approaches to understand the mechanistic links among the processes that accompany and/or lead to aging.
-
Microbiome
Understanding of the human microbiome has grown exponentially over the past decade. ISB researchers are unlocking the mysteries of the human microbiome and translating our scientific knowledge into therapies for a number of complex diseases.
-
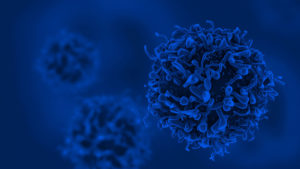
Cancers
Cancer research poses many interconnected complexities related to early detection, stratification and treatment. The cross-disciplinary holistic, integrative nature of systems biology makes ISB researchers well suited to tackle the complexity of these diseases.
-

Brain Health
ISB, along with a collaborative network of partners, is pioneering a multimodal approach that combines personal data, lifestyle factors, cognitive training and systems medicine — and is rigorously testing these new approaches in clinical trials. This approach is critical to prevent, slow, and even reverse many neurological conditions before they become irreversible.
-
Scientific Wellness
Scientific Wellness embodies P4 medicine (predictive, preventive, personalized and participatory), and is being applied to help individuals improve their health. The platform provides new approaches for drug target discovery, and is driving profound economic, policy and social change.
-

Pregnancy Health
By using a systems approach to pregnancy health, ISB is gaining a more comprehensive understanding of pathological changes that occur in pregnancy, which will identify women most at risk of developing pregnancy complications.
-

Environment
Achieving environmental sustainability is one of civilization’s historic challenges. ISB uses systems science to improve the understanding of the interactions of microbes and ecosystems and create a new generation of sustainable tools and strategies that can be deployed to address this challenge.
Privacy Overview
This website uses cookies so that we can provide you with the best user experience possible. Cookie information is stored in your browser and performs functions such as recognising you when you return to our website and helping our team to understand which sections of the website you find most interesting and useful.



 isbscience.org/research/
isbscience.org/research/