Systems biology always looks for answers in more than one place.
Systems biology focuses on untangling molecular, genetic, and environmental interactions within biological systems in order to understand and predict behavior in living organisms.

In a well-known story, six blind men come across an elephant for the first time. Each one touches a different part of the elephant to figure out what it is. One touches the trunk and thinks the elephant is like a thick snake. Another touches a leg and believes it’s like a tree, and so on. Each blind man makes a guess, but they will never fully understand the elephant until they share their information. Systems biology addresses this challenge by taking a big-picture approach to studying complex biological systems, combining different perspectives to understand the whole system. Image credit: ISB
What is Systems Biology?
Systems biology is an interdisciplinary approach to science that aims to understand how biological components—such as genes, proteins, and cells—interact and function together as a system. Systems biology integrates various fields of study, including genomics, proteomics, metabolomics, and other “omics” areas (known as multiomics), to construct comprehensive predictive models and simulate the behavior of biological systems under various conditions.
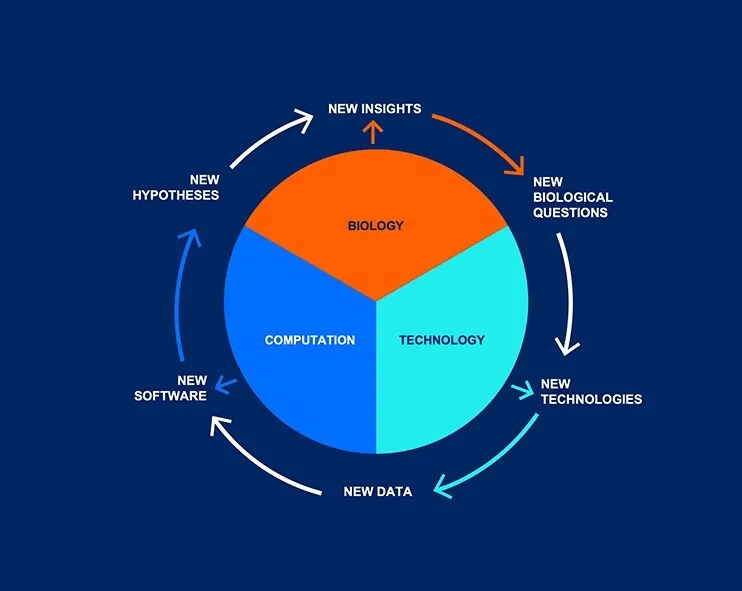
The Innovation Engine
Solving challenging biological problems requires the development of new technologies for exploring previously unknown properties of biological systems. These technologies represent new dimensions of data space. Each new data type requires novel analytical tools, creating a virtuous cycle where biology drives technology, and technology drives computation. New biological insights emerge from each iteration of this cycle, generating novel technologies and computational tools. Finally, we disseminate our innovations through open-access platforms, spinout companies, and beyond, ensuring our advancements benefit the broader scientific community. We call this the “innovation engine.”

A Network of Networks
Our bodies are composed of many networks of molecular and cellular interactions that integrate and communicate across multiple scales. From our genome to the molecules and cells that form our organs, and extending out to our interactions within the world, we are fundamentally a network of networks. Systems biology examines these networks, integrating behaviors at various levels to formulate hypotheses about biological functions. This approach provides insights into dynamic biological changes, enhancing our understanding of life’s complexity and informing innovative solutions in health and disease.
Data Integration
Systems biology relies on data integration, which allows researchers to combine and analyze diverse types of biological data – from multiomic data to electronic health records to quantified self-data that includes diet and fitness – allowing us to gain comprehensive insights into complex biological systems. Integrating these diverse data sets leads to the development of more accurate computational models and predictive tools, driving innovation in research and healthcare. This approach enhances our understanding of biological functions and disease mechanisms, paving the way for advancements in personalized medicine and targeted therapies.
Cross-Disciplinary Teams
Biological complexity in the age of “big data” requires multi-disciplinary, diverse teams to come together. One of the Institute for Systems Biology’s research labs, for example, includes molecular biologists, microbiologists, geneticists, engineers, oceanographers, and even an astrophysicist. These complementary skills empower our researchers to understand biological and environmental challenges from different perspectives so they can achieve shareable insights more quickly.
Multiomics
Multiomics, a now-ubiquitous term among scientists, describes the integration of information across various “-omes” of a biological system, such as the genome, transcriptome (mRNAs), proteome (proteins), microbiome, epigenome, metabolome, and phenome. Through examination of these interconnected layers of biological information, multiomics provides a deeper understanding of health and disease, driving advancements in research and healthcare.
Predictive Modeling
By enabling the simulation and analysis of complex biological interactions, predictive models allow a deeper understanding of life’s complexities and support the development of innovative solutions to biological and medical challenges. One exciting concept in this process is the digital twin, where a virtual replica of a biological entity (e.g., a patient) uses real-world data to run computer simulations under a variety of conditions to predict how a patient will respond to different treatments.
Help us realize a new vision of health.
The Institute for Systems Biology (ISB) is a nonprofit scientific research organization located in Seattle. We believe in the power of science to transform health. If you are committed to a better future, ISB is here to make that a reality.